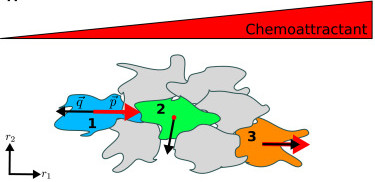
A collaborative paper with Professor Andrew Mugler’s group in Physics is published in Biophysical Journal, and the result is highlighted at the Journal website (Video).
Collective Chemotaxis through Noisy Multicellular Gradient Sensing
Julien Varennes, Bumsoo Han, Andrew Mugler
Abstract
Collective cell migration in response to a chemical cue occurs in many biological processes such as morphogenesis and cancer metastasis. Clusters of migratory cells in these systems are capable of responding to gradients of <1% difference in chemical concentration across a cell length. Multicellular systems are extremely sensitive to their environment, and although the limits to multicellular sensing are becoming known, how this information leads to coherent migration remains poorly understood. We develop a computational model of multicellular sensing and migration in which groups of cells collectively measure noisy chemical gradients. The output of the sensing process is coupled to the polarization of individual cells to model migratory behavior. Through the use of numerical simulations, we find that larger clusters of cells detect the gradient direction with higher precision and thus achieve stronger polarization bias, but larger clusters also induce more drag on collective motion. The trade-off between these two effects leads to an optimal cluster size for most efficient migration. We discuss how our model could be validated using simple, phenomenological experiments.